With the advent of metagenomic studies, scientists need to understand the terminology used in them. Much like metagenomics itself, the vocabulary used contains words from multiple disciplines such as ecology, bioinformatics, and microbiology. However, these terms may have slightly different meaning and nuances when used in metagenomics studies. To help clear up the usage of some of these terms and provide a little back ground on them, I have complied a guide that breaks down the meaning and associated nuances for some of the most frequently used terms in metagenomics studies.
Microbiome- All the microbial (bacterial, fungal, viral, eukaryote, or a combination of any of these) DNA in an environment, community, or ecosystem. This data is usually obtained by use of high throughput sequencing. This term functions as a microbial census of “who is there?” Results can be obtained through 16S, 18S, and ITS markers (which will be defined more fully later.) Because microbiomes are composed of DNA found through sequencing, just because an organisms’ DNA is part of the microbiome doesn’t mean that organism is alive within the community.
Microbiota- All the microbes (bacterial, fungal, viral, eukaryote, or a combination of any of these) in an environment, community, or ecosystem. This is subtly different from the microbiome because it describes all of the organisms present, not just the DNA found. These microbes are the ones alive and functioning in a particular environment, community, or ecosystem and can also encompass all microbes, bacteria, fungi, virus, or some eukaryotes.
Metagenomics- The genetic analysis of microorganisms in conjunction with relevant data. These studies extract total DNA from a sample and collect information about that sample that can be useful in analysis. These studies often revolve around “who is there” and/or “what are they doing”. By characterizing a microbial community and learning how that microbial community functions, or shifts can provide insight into microbial based intervention and a greater understanding of microbial ecology.
WGS- This is an acronym that stands for “Whole Genome Shotgun Sequencing”, sometime this is just referred to as “Shotgun”. WGS is method of determining the entire sequence of a genome (for example a bacterial genome) by the use of sequencing fragments of the genome and piecing them back together. The genome is broken into small segments which are then sequenced and put back together with a reference genome or de novo (done without a reference) by utilizing to overlapping portions of the segment. The FDA is employing this kind of methodology in GenomeTrackr to aid in outbreak tracking and identification.
OTU- This term stands for “Operational Taxonomic Unit”. It is an arbitrary designation that is used to denote a distinct sequence and is synonymous to a species. However, when processing data obtained from next gen sequencing, it cannot be given a taxonomic identity until it has been compared to a reference set of sequences with determined taxonomy. It is very common to BLAST OTUs to obtain the information.
16S- This is a ribosomal subunit found in all bacteria that is used to determine the microbiome of a sample. The sequence is highly conserved but has enough difference to reveal taxonomy when sequence. IT is also present in all bacteria and microbes often contain multiple copies of this subunit in their genome. All of these features make it an ideal target for microbiome studies for bacteria. The regions targeted in high throughput next generation sequencing are the hypervariable regions (V1-V9) and usually a 100-200 bp fragment of these regions are used for sequencing.
18S- This is a ribosomal subunit found in all eukaryotes that is used to determine the microbiome of a sample. The 16S analogue for eukaryotes, the 18S sequence is highly conserved, specific to eukaryotes, and an ideal target for microbiome studies.
ITS- The ITS (Internal Transcribed Spacer) is found within all fungi and is used to determine the microbiome of a sample. Similar to 16S rRNA and 18S rRNA, the ITS region is common to all fungi and has enough variation to delineate taxonomy. Although this has been changing, historically fungal diversity has not been well studies so the best region with the ITS to target is still debated.
Alpha Diversity- This term applies to the diversity within a single sample. The term is often applied in ecology to describe the number and distribution of species. The same principals are applied to the microbial ecology, the microbiome or microbiota, of a sample.
Beta Diversity- This term describes the diversity and difference between samples. It allows for the comparison of one sample to another based on diversity. It is another term with roots in ecology that has been applied to the microbial ecology world as well.
Shannon Diversity- This is a measure of alpha diversity within a community. It accounts for both the abundance and evenness of species present across the community. However, the function can be applied at any level of microbial taxonomy (family, genus, species, OTU, etc) depending on how the data is being analyzed.
UniFrac- This is a distance metric used to compare the similarity of biological community samples. The analysis can be weighted, which is a quantitative measure, or unweighted, which is a qualitative measure. This comparison is achieved by using a phylogenetic tree model where taxa that are common to both samples being compared are considered “shared” and the distance between two samples is calculated by comparing the shared and unshared branches.
QIIME- This stands for “Quantitative Insights Into Microbial Ecology” and is an open-access platform for data analysis of microbiomes and metagenomics data sets. QIIME use’s command line based python coding in a terminal that can be installed on a computer for free. Every step of microbiome and metagenomic data analysis, from creating contigs to assigning taxonomy and creating PCAs, can be performed on QIIME. The software allows for complete control of quality cut offs and even allows you to compare OTUs against the database of your choice.
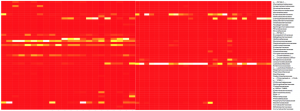
MG-RAST- The acronym stands for “Metagenomic Rapid Annotations using Subsystems Technology”. This is another open-access system used for the analysis of microbiomes and metagenomics. The program is web based and will allow other to access and manipulate data that is submitted on the website. Aligned and merged read, or contigs, are submitted with metadata and within a week or two results are obtained. While there is less control over analysis in this pipeline, all the data is processed in the same way and it allows for comparison of any of the submitted files on the website to one another. It can create heat maps, taxonomy charts and a written summary on the quality of data.
GENIUS- This is a software program that goes far beyond metagenomics and microbiome analysis. This software allows the processing of many different kinds of sequencing data from creating phylogenetic trees to merging reads. They do have a metagenomics package that can create interesting and interactive diagrams based on the taxonomic composition of samples.
Phinch- This is another web-based open-access tool for analyzing microbiome and metagenomics data. It allows the user to upload a biome file and creates several kinds of diagrams (bubble charts, graphs donut comparisons, etc) to aid in data visualization. However, it only works for the older format (1.8) of QIIME biome files so make sure to run your data through that if you plan to utilize this tool.
